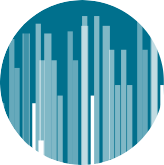
The UMGC’s latest sequencing instrument, the AVITI System by Element Biosciences, is a short-read, mid-throughput sequencer that delivers a lower cost per GB without the need to scale throughput.
This instrument features two flow cells, each operating independently, to give the UMGC greater sequencing flexibility while maintaining the option to deliver up to 2 billion reads of combined sequencing output.* The AVITI delivers the most accurate short-read sequencing available today, with most reads scoring above Q40.*
Element achieves higher sequencing accuracy via a novel sequencing technology, termed Avidity Sequencing, that optimizes nucleotide incorporation and identification steps while reducing the concentration of reagents.
The AVITI readily adapts to a variety of applications, offering methods that scale from amplicons to whole genomes. With outstanding data quality and compatibility with existing library formats, the AVITI will be of particular interest to single-cell and spatial researchers, as well as researchers who require ultra-high-quality reads. Learn more about the Element AVITI system.
Highlighted applications
- Single-cell and bulk RNA sequencing
- Whole genome metagenomic sequencing
- Spatialomic sequencing
- Ultra-low pass sequencing (Skim-seq)
- Whole genome human and mouse sequencing (~3 samples @ 30X coverage)
- Element LoopSeq 16S and amplicon sequencing (synthetic long reads)
Output
The AVITI flow cells come in low, medium, and high outputs with 150, 300, and 600-cycle sequencing kits. The kits use the newer and faster Avidity Cloudbreak chemistry that enables run configurations to reach 1 billion reads or more with a maximum sequencing time of 50-60 hours.**
Benefits
Element’s proprietary innovations in chemistry and instrumentation have significantly reduced sequencing costs for researchers while simultaneously delivering unprecedented sequencing data quality. With the introduction of the AVITI System, UMN researchers will now be able to purchase sequencing with an average of one error per 10,000 base pairs (Q40) and at 90% of bases scoring Q30*.
Library Options
Element has multiple options to streamline library prep for sequencing on the AVITI:
- Adept Workflow for adapted libraries - circularizes linear libraries prepared with a compatible third-party library prep kit (i.e. Illumina) to allow researchers to continue using the same library prep and analysis tools. The UMGC anticipates this to be the most commonly requested workflow.
- Elevate Workflow for native prep - prepares dual-indexed linear libraries from genomic DNA input and libraries are circularized on the flow cell as part of the sequencing run. Include the choice of PCR-free or PCR-plus.
- LoopSeq for AVITI - prepares libraries for long-read sequencing on the short-read Element AVITI System. Available for 16S and amplicon sequencing.
UMN Rates
High Output Sequencing Rates Summary
| 150-cycle (75 PE) | Savings Tier | Speed Tier | Priority Tier |
|---|---|---|---|
| Medium run. 500 M reads/lane. | $781.72 | $940.12 | $1,172.57 |
| Turnaround time. | 5 - 11 days | 2 - 5 days | 2 - 3 days |
| High run. 1 B reads/lane. | $1,131.36 | $1,347.96 | $1,667.71 |
| Turnaround time. | 5 - 11 days | 2 - 5 days | 2 - 3 days |
| 300-cycle (150 PE) | Savings Tier | Speed Tier | Priority Tier |
|---|---|---|---|
| Low run. 250 M reads/lane. | $900.67 | $1,078.87 | $1,341.02 |
| Turnaround time. | 6 - 13 days | 3 - 7 days | 3 - 5 days |
| Medium run. 500 M reads/lane. | $1,130.16 | $1,266.70 | $1,506.29 |
| Turnaround time. | 6 - 13 days | 3 - 7 days | 3 - 5 days |
| High run. 1B reads/lane. | $1,417.33 | $1,757.48 | $1,992.99 |
| Turnaround time. | 6 - 13 days | 3 - 7 days | 3 - 5 days |
| 600-cycle (300 PE) | Savings Tier | Speed Tier | Priority Tier |
|---|---|---|---|
| Stowaway. 25 M reads. | $593.38 | N/A | N/A |
| Turnaround time. | 2 - 4 weeks | N/A | N/A |
| Medium run. 100 M reads/lane. | $1,431.75 | $1,698.35 | $2,093.10 |
| Turnaround time. | 7 - 17 days | 4 - 10 days | 4 - 7 days |
| High run. 300 M reads/lane. | $1,979.64 | $2,337.44 | $2,868.99 |
| Turnaround time. | 7 - 17 days | 4 - 10 days | 4 - 7 days |
Notes
1. Output: Custom libraries and libraries that have not been tested using Adept conversion may not achieve the above output without some initial qualification. Current tested libraries: Illumina NexteraXT, Illumina DNA, SMARTer Stranded Total RNA-Seq Kit v2 - Pico Input Mammalian, 10X 3' and 5' GEX. See compatible kits for the Element Adept Library Compatibility Workflow. Contact next-gen@umn.edu if you have an interest in a kit not contained in this list.
2. Tiers: Turnaround time (TAT) is in business days. TAT is for sequencing only and does not factor in the TAT for library prep or other companion services. Availability for the Priority Tier is limited; contact next-gen@umn.edu to see if this is a current option. More information about Pricing Tiers.
External Rates
High Output Sequencing Rates Summary
| 150-cycle (75 PE) | Savings Tier | Speed Tier | Priority Tier |
|---|---|---|---|
| Medium run. 500 M reads/lane. | $952.62 | $1,119.72 | $1,367.21 |
| Turnaround time. | 5 - 11 days | 2 - 5 days | 2 - 3 days |
| High run. 1 B M reads/lane. | $1,381.77 | $1,607.07 | $1,941.86 |
| Turnaround time. | 5 - 11 days | 2 - 5 days | 2 - 3 days |
| 300-cycle (150 PE) | Savings Tier | Speed Tier | Priority Tier |
|---|---|---|---|
| Low run. 250 M reads/lane. | $1,098.61 | $1,285.51 | $1,562.70 |
| Turnaround time. | 6 - 13 days | 3 - 7 days | 3 - 5 days |
| Medium run. 500 M reads/lane. | $1,380.29 | $1,525.53 | $1,780.16 |
| Turnaround time. | 6 - 13 days | 3 - 7 days | 3 - 5 days |
| High run. 1 B reads/lane. | $1,682.54 | $2,045.95 | $2,306.31 |
| Turnaround time. | 6 - 13 days | 3 - 7 days | 3 - 5 days |
| 600-cycle (300 PE) | Savings Tier | Speed Tier | Priority Tier |
|---|---|---|---|
| Stowaway. 25 M reads. | $673.99 | N/A | N/A |
| Turnaround time. | 2 - 4 weeks | N/A | N/A |
| Medium run. 100 M reads/lane. | $1,750.46 | $2,025.76 | $2,435.55 |
| Turnaround time. | 7 - 17 days | 4 - 10 days | 4 - 7 days |
| High run. 300 M reads/lane. | $2,422.93 | $2,789.43 | $3,336.02 |
| Turnaround time. | 7 - 17 days | 4 - 10 days | 4 - 7 days |
Notes
1. Output: Custom libraries and libraries that have not been tested using Adept conversion may not achieve the above output without some initial qualification. Current tested libraries: Illumina NexteraXT, Illumina DNA, SMARTer Stranded Total RNA-Seq Kit v2 - Pico Input Mammalian, 10X 3' and 5' GEX. See compatible kits for the Element Adept Library Compatibility Workflow. Contact next-gen@umn.edu if you have an interest in a kit not contained in this list.
2. Tiers: Turnaround time (TAT) is in business days. TAT is for sequencing only and does not factor in the TAT for library prep or other companion services. Availability for the Priority Tier is limited; contact next-gen@umn.edu to see if this is a current option. More information about Pricing Tiers.
Submission
How to order
Download the appropriate submission form for submitting samples or pre-made libraries. Once the form has been completed, please submit the submission by emailing next-gen@umn.edu. Samples can be dropped off at one of our campus locations
1-210 Cancer & Cardiovascular Research Building (Minneapolis campus)
20 Snyder Hall (St. Paul campus)
Samples can be shipped to the following address:
Please send the tracking information to next-gen@umn.edu.
UMN Genomics Center
ATTN: NGS Staff
3510 Hopkins Place N.
Building 4 Suite W402
Oakdale, MN 55128
612-625-7736
Deliverables
Data Release
There are four options for transferring data from the UMGC to clients: 1) delivery to the Minnesota Supercomputing Institute’s (MSI) high-performance file system, 2) download from a secure website, 3) download with Globus, or 4) shipment on an external hard drive. Please indicate your data delivery preference when placing an order for sequencing.
1. MSI storage
Internal clients have their data released to MSI's Shared User Resource Facility Storage (SURFS). Delivered data will be located in the "data_delivery" folder in your group's folder on MSI's primary filesystem (home/GROUP/data_delivery/umgc). MSI does not charge for SURFS storage costs, but files expire and are removed one year after they've been delivered. Files should be copied to other MSI storage locations such as Tier2, Tier3, or your group's "shared" folder before they expire.
2. Web download
Internal clients that opt-out of MSI storage and external clients can download their data from a secure website using either a web browser or a command-line download tool, complete instructions are provided in an email from the UMGC. The client’s data is available for download for 30 days, after which the data will be removed from the data download website and the client takes responsibility for storing the data.
3. Globus
Internal and External clients can use Globus to download their data. This is the recommended method for external clients to download large datasets.
4. Hard drive
External clients may have data shipped on a hard drive purchased by the UMGC and invoiced to the client at a cost of $250 per hard drive.
Data Recovery
The UMGC archives most customer data for a year and some datasets are retained for 5 years or more. If you need a dataset re-delivered email a request to next-gen@umn.edu to initiate data recovery. The UMGC does not provide any guarantee that data can be successfully recovered from the archive.
FAQs
Do you accept client-made libraries?
Yes, please consult with UMGC staff before submitting client-made libraries by emailing next-gen@umn.edu.
We recommend that pooling be done at UMGC as PicoGreen concentrations are used for balancing libraries. However, clients may choose to pool themselves.
Client-made libraries will have the following services performed on them before proceeding with sequencing.
- PicoGreen (concentration of library)
- DNA Agilent High Sensitivity chip (size of library)
- KAPA QC (functionality of library)
Where should samples be shipped?
University of Minnesota Genomics Center
1475 Gortner Ave.
28 Snyder Hall
St. Paul, MN 55108
612-625-7736